Regression and Clustering
In the previous chapter we used a K-Nearest Neighbors classifier with scikit-learn. Here we expand our coverage of “classic” machine learning with two more fundamental techniques: regression for predicting continuous values, and clustering for discovering groups in data without labeled targets.
The requirements for this chapter are:
1 uv pip install scikit-learn pandas numpy
The examples for this chapter are in the directory source-code/regression_and_clustering.
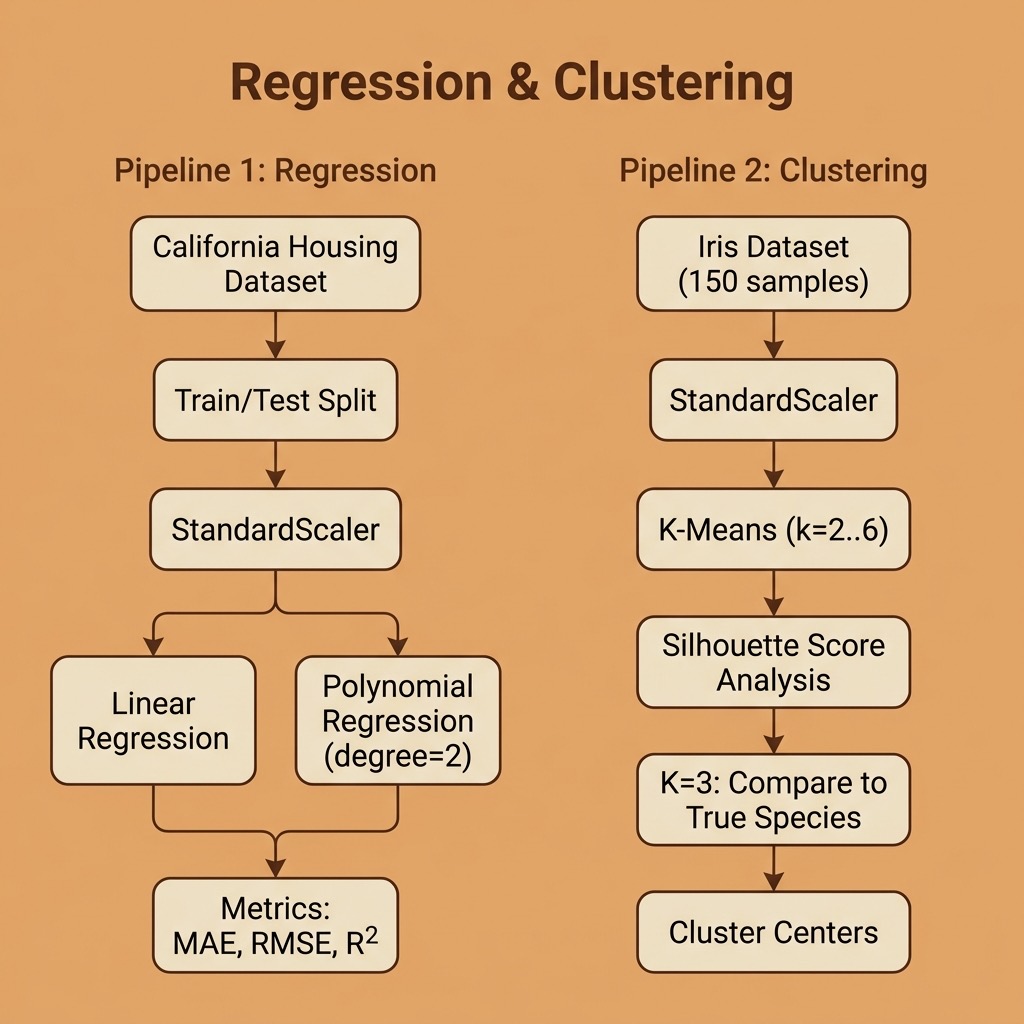
Regression: Predicting Housing Prices
Regression is a type of supervised machine learning where the goal is to predict a continuous numerical value rather than a class label. While classification asks “which category?”, regression asks “how much?” or “how many?”
Common regression algorithms include:
- Linear Regression: models the relationship between features and target as a straight line (or hyperplane in multiple dimensions).
- Polynomial Regression: extends linear regression by adding polynomial terms, allowing the model to capture non-linear relationships.
- Ridge and Lasso Regression: variants of linear regression that add regularization to prevent overfitting.
We will use the California Housing dataset, which is built into scikit-learn. It contains 20,640 samples with 8 features describing California census block groups (median income, average number of rooms, location, etc.), with the target being the median house value in units of $100,000s.
Linear Regression
Our first example fits a linear regression model to all 8 features:
1 import numpy as np
2 import pandas as pd
3 from sklearn.datasets import fetch_california_housing
4 from sklearn.linear_model import LinearRegression
5 from sklearn.preprocessing import PolynomialFeatures, StandardScaler
6 from sklearn.metrics import mean_absolute_error, mean_squared_error, r2_score
7 from sklearn.model_selection import train_test_split
8
9
10 def load_housing_data():
11 """Load the California Housing dataset into a DataFrame."""
12 housing = fetch_california_housing()
13 df = pd.DataFrame(housing.data, columns=housing.feature_names)
14 df["MedHouseVal"] = housing.target
15 return df
We use train_test_split to hold out 20% of the data for evaluation, scale the features with StandardScaler, and train the model:
1 df = load_housing_data()
2 X = df.drop("MedHouseVal", axis=1)
3 y = df["MedHouseVal"]
4
5 X_train, X_test, y_train, y_test = train_test_split(
6 X, y, test_size=0.2, random_state=42
7 )
8
9 scaler = StandardScaler()
10 X_train_scaled = scaler.fit_transform(X_train)
11 X_test_scaled = scaler.transform(X_test)
12
13 model = LinearRegression()
14 model.fit(X_train_scaled, y_train)
15 y_pred = model.predict(X_test_scaled)
To evaluate a regression model, we use different metrics than classification:
- MAE (Mean Absolute Error): average magnitude of prediction errors in the same units as the target.
- RMSE (Root Mean Squared Error): like MAE but penalizes larger errors more heavily.
- R² (R-squared): the proportion of variance in the target explained by the model (1.0 = perfect, 0.0 = no better than predicting the mean).
Here is the output of running the full regression.py script:
1 $ python regression.py
2 Dataset shape: (20640, 9)
3 Target range: 0.15 - 5.00 (units: $100,000s)
4
5 === Linear Regression Results ===
6 MAE: 0.5332
7 RMSE: 0.7456
8 R²: 0.5758
9
10 Feature coefficients (scaled):
11 MedInc +0.8544
12 HouseAge +0.1225
13 AveRooms -0.2944
14 AveBedrms +0.3393
15 Population -0.0023
16 AveOccup -0.0408
17 Latitude -0.8969
18 Longitude -0.8698
Because we scaled the features, the coefficients tell us the relative importance of each feature. MedInc (median income) has the largest positive coefficient, confirming that income is the strongest predictor of housing prices. The geographic features (Latitude and Longitude) are also very influential — location matters!
An R² of 0.58 means our simple linear model explains about 58% of the variance in housing prices. Not bad for a single line of model code, but there is room for improvement.
Polynomial Regression
We can capture non-linear relationships by creating polynomial features. For interpretability, we use just two features (MedInc and AveRooms) and generate all degree-2 combinations:
1 poly = PolynomialFeatures(degree=2, include_bias=False)
2 X_train_poly = poly.fit_transform(X_train_scaled)
3 X_test_poly = poly.transform(X_test_scaled)
4
5 model = LinearRegression()
6 model.fit(X_train_poly, y_train)
The PolynomialFeatures transformer with degree=2 creates five features from our original two: x0, x1, x0², x0·x1, and x1². The interaction term x0·x1 lets the model learn that the relationship between income and price depends on the number of rooms.
1 === Polynomial Regression (degree=2, 2 features) ===
2 MAE: 0.6163
3 RMSE: 0.8402
4 R²: 0.4613
5
6 Polynomial feature names: ['x0', 'x1', 'x0^2', 'x0 x1', 'x1^2']
With only 2 features, the polynomial model achieves R²=0.46 compared to 0.58 with all 8 features linearly. This illustrates an important tradeoff: polynomial features add expressive power but using too few input features limits the information available to the model.
Clustering: Discovering Groups in Data
Clustering is an unsupervised learning technique — unlike classification and regression, we do not provide labeled targets. Instead, the algorithm discovers natural groupings in the data.
K-Means is the most widely used clustering algorithm. It works by:
- Choosing K initial cluster centers (centroids).
- Assigning each data point to the nearest centroid.
- Recomputing each centroid as the mean of its assigned points.
- Repeating steps 2-3 until convergence.
We will use the classic Iris dataset (150 samples, 4 features measuring sepal and petal dimensions, with 3 known species). This lets us evaluate how well K-Means recovers the true species groupings.
Choosing K with the Silhouette Score
A common question is: how many clusters should we use? The silhouette score measures how similar each point is to its own cluster compared to other clusters (ranging from -1 to +1, higher is better):
1 from sklearn.cluster import KMeans
2 from sklearn.preprocessing import StandardScaler
3 from sklearn.metrics import silhouette_score
4
5 scaler = StandardScaler()
6 X_scaled = scaler.fit_transform(X)
7
8 for k in range(2, 7):
9 km = KMeans(n_clusters=k, random_state=42, n_init=10)
10 labels = km.fit_predict(X_scaled)
11 score = silhouette_score(X_scaled, labels)
12 print(f" k={k}: silhouette score = {score:.4f}")
Output from running clustering.py:
1 $ python clustering.py
2 Dataset: 150 samples, 4 features
3 Features: ['sepal length (cm)', 'sepal width (cm)',
4 'petal length (cm)', 'petal width (cm)']
5 Known species: ['setosa', 'versicolor', 'virginica']
6
7 === Silhouette Scores for Different k Values ===
8 k=2: silhouette score = 0.5818
9 k=3: silhouette score = 0.4599
10 k=4: silhouette score = 0.3869
11 k=5: silhouette score = 0.3459
12 k=6: silhouette score = 0.3171
The silhouette score is highest at k=2, which suggests that the data most naturally splits into two groups. This makes biological sense: setosa is very distinct from the other two species, while versicolor and virginica overlap considerably.
Comparing Clusters to True Labels
Since we happen to know the true species labels, we can use a cross-tabulation to see how well K-Means (with k=3) recovers them:
1 === K-Means (k=3) Cluster vs. True Species ===
2 Cluster 0 1 2
3 Species
4 setosa 0 50 0
5 versicolor 39 0 11
6 virginica 14 0 36
K-Means perfectly separates all 50 setosa samples into cluster 1. However, it struggles to distinguish versicolor from virginica — 14 virginica samples are assigned to the versicolor-dominant cluster 0, and 11 versicolor samples end up in the virginica-dominant cluster 2. This confirms what the silhouette scores suggested.
The cluster centers (transformed back to the original scale) reveal the “typical” flower in each group:
1 === Cluster Centers (original scale) ===
2 sepal length sepal width petal length petal width
3 Cluster
4 0 5.80 2.67 4.37 1.41
5 1 5.01 3.43 1.46 0.25
6 2 6.78 3.10 5.51 1.97
Cluster 1 (setosa) has much smaller petals, while cluster 2 (mostly virginica) has the largest sepals and petals.
Regression and Clustering Wrap-up
In this chapter we covered two fundamental machine learning techniques beyond classification:
- Regression lets us predict continuous values. We saw that even simple linear regression can be informative, and that feature coefficients reveal which inputs drive predictions.
- Clustering discovers structure in unlabeled data. We used silhouette scores to evaluate cluster quality and cross-tabulations to compare clusters against known labels.
Together with the classification example in the previous chapter, you now have hands-on experience with the three core tasks of classic machine learning. In the next chapter we will explore the data preparation work that comes before training any model.