Chapter 7 - fMRI basics: Processing stages, terminology, and data structure
fMRI basics
In this chapter, we’ll talk about analysis of functional magnetic resonance imaging, or fMRI, data. We’ll start with some nomenclature, talk about data types and structures, and end with a little bit about fMRI data analysis goals. fMRI is a noninvasive technique for studying brain activity. By noninvasive, we mean that it generates pictures of the inside of your head without using any implants or injections. There are also no known side effects of being scanned frequently with fMRI. For example, one of our colleagues scanned himself nearly 100 times with no apparent adverse effects. Scans are now also routinely performed on both infants and children.
A single session in the scanner allows researchers to collect many image types, both anatomical (or `structural’ MRI) and functional (related to dynamic brain activity changes). Here, we are concerned particularly with functional imaging.
During the course of an fMRI experiment a series of brain images are acquired, often while the subject performs a task set. Those ‘tasks’ can include cognitive paradigms, viewing, hearing, feeling, tasting, or smelling various stimuli. An increasingly popular ‘task’ is simply lying in the scanner doing nothing; there is now a whole subfield devoted to ‘resting-state’ fMRI.
Processing and analysis stages
Figure 7.1 shows a basic flowchart for a typical fMRI experiment. Throughout this book, we’re going to keep coming back to this flowchart to unpack each of the basic data processing and analysis steps in more detail. All studies begin with experimental design, which is perhaps the single most crucial factor in determining how well the experiment will go. There are a number of design principles rooted in statistics which apply to all design types, neuroimaging or otherwise. Other principles are specific to fMRI experiments and relate to the properties of fMRI data and their specific analyses.

Data acquisition is the next step after experimental design, followed by reconstruction of the data into images. Researchers perform a series of preprocessing steps before statistical analysis. These deal with anatomical alignment of the various image types (coregistration), timing issues (slice-timing correction), head movement (motion correction), and image transformation onto a standard anatomical reference space (spatial normalization or warping). Researchers also commonly perform artifact-mitigation procedures and physiological noise correction. After preprocessing, the images are ready for statistical analysis. Analyses can test task- or outcome-related brain activity, assess functional connectivity, and develop multivariate predictive models designed to correlate optimally with experimental variables or outcomes.
Acquisition
We acquire MRI and fMRI data by applying radiofrequency (RF) pulses to the brain. These pulses perturb the magnetic spins of the protons of hydrogen atoms (mostly in water molecules) so they give off energy with particular spatiotemporal characteristics. The RF antenna reads off this signal, which is then used to reconstruct images. During data acquisition, magnetic gradient coils are applied in particular patterns so signals from different spatial locations in the image are given particular characteristics, which enables accurate spatial reconstruction. The pulse sequence, the software that runs the RF antenna and gradient coils which acquire the signal, determines what type of data the process acquires - including whether the image is structural or functional. This will all be covered in greater detail in a later chapter.
It’s useful to know some basic terminology related to MR image acquisition. Figure 7.2 shows some of the basics. The bounding box that defines the image acquisition volume depends on the field of view (the slice dimensions), the number of slices, and the slice thickness. Data are sampled within small cubic volumes called `voxels’ or volumetric pixels. Voxel size depends on slice thickness and in-plane matrix size, which is the number of grid elements on which each slice’s data are sampled. The field of view divided by the matrix size is the in-plane resolution, measured in mm. Thus in-plane resolution and slice thickness determine voxel size. Researchers typically desire isotropic voxels, which are the same dimension on all sides, though unequal sizes work as well. A typical size is  voxels; this is close to optimal for many purposes when using a 3-Tesla scanner.
voxels; this is close to optimal for many purposes when using a 3-Tesla scanner.

Designing an fMRI study requires a series of tradeoffs given the study’s particular goals. One fundamental tradeoff is between spatial and temporal resolution. You can either collect data with high spatial resolution or collect data fast, but you can’t do both. Spatial resolution defines our ability to distinguish how an image changes across different spatial locations and thus also our ability to extract location-coded information about brain states and behaviors. Both the voxel size and the image’s underlying smoothness (blurriness), which depends on the main magnetic field’s strength and gradients and on underlying physiological limitations (because most of our signal is blood flow-related), determine spatial resolution. Temporal resolution determines our ability to separate brain events in time. Both the TR and the hemodynamic response to neural and/or glial events’ time course (more on this is below) determine this.
Image orientation and dimensions
Understanding and interpreting which part of the brain one is viewing requires some practice. The brain is a complex three-dimensional structure with many curved Ã’CÓ shape sub-structures that wrap around the brain’s center, the thalamus. It is typical to show neuroimaging results on anatomical brain slices. Figure 7.3 provides a basic orientation to those slices and their spatial relation to the overall head and brain surface. Each of the three dimensions of brain space has a special name. The left-to-right dimension is conventionally the 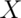 direction in standard brain coordinate space. The back-to-front dimension is the
direction in standard brain coordinate space. The back-to-front dimension is the 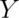 dimension which ranges from posterior at the brain’s back to anterior at the front. Sometimes anterior is also called rostral, which means ‘toward the head’, and posterior is called caudal, ‘toward the tail’. The bottom-to-top dimension is the
dimension which ranges from posterior at the brain’s back to anterior at the front. Sometimes anterior is also called rostral, which means ‘toward the head’, and posterior is called caudal, ‘toward the tail’. The bottom-to-top dimension is the 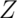 dimension which ranges from inferior to superior locations. These locations are sometimes also called ventral (‘towards the belly’) and dorsal (‘towards the back’).
dimension which ranges from inferior to superior locations. These locations are sometimes also called ventral (‘towards the belly’) and dorsal (‘towards the back’).

Researchers typically report locations along these dimensions in ![[x, y, z]](/preview_site_images/principlesoffmri/leanpub_equation_11.png) coordinate triplets with
coordinate triplets with 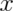 ,
, 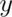 , and
, and 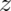 values indicating distances in millimeter units relative to a zero point. The
values indicating distances in millimeter units relative to a zero point. The ![[0, 0, 0]](/preview_site_images/principlesoffmri/leanpub_equation_15.png) point is, by convention, the anterior commissure, a small white-matter bundle which connects the brain’s two hemispheres. Figure 7.4 shows this point.
point is, by convention, the anterior commissure, a small white-matter bundle which connects the brain’s two hemispheres. Figure 7.4 shows this point.

In the brainstem, some of the dimension names are not very intuitive because they describe the dimensions as in an animal which walks on four legs with the spinal cord toward the rear (caudal) and the midbrain, which lies just below the thalamus, at the rostral end. Thus, the part of the brainstem toward the back of the head is the dorsal brainstem and the part toward the front is the ventral brainstem. Figure 7.5 shows these directions and some of the most important structures’ locations.

The sections of the brain that are typically used to display neuroimaging results have particular names too. Figure 7.3 shows these. Coronal slices are sections that span the left-right and inferior-superior dimensions at one location from front to back. Sagittal slices span the front-to-back and inferior-superior dimensions at one location from left to right. And axial or *horizontal * slices span the left-right and front-to-back dimensions at one location from inferior to superior.
fMRI time series
Functional images (also called 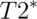 -weighted images) have lower spatial resolution than structural images. That is, they’re much blurrier than their structural counterparts. However, we can measure many of them, so they have higher temporal resolution and we can use them to relate signal changes to experimental manipulations or other outcomes that vary from second to second.
-weighted images) have lower spatial resolution than structural images. That is, they’re much blurrier than their structural counterparts. However, we can measure many of them, so they have higher temporal resolution and we can use them to relate signal changes to experimental manipulations or other outcomes that vary from second to second.
One participant’s fMRI dataset contains a time series of 3-D images, or `volumes’, shown in Figure 7.6. The volumes often cover the entire brain but can also cover just one brain tissue section or slab at a higher spatial resolution. The data for each volume are usually acquired slice-by-slice; after completing one volume, the scanner moves on to the next image. As they are collected, the data are sampled onto a rigid voxel grid.

It is not uncommon for each volume to contain 100,000 or more voxels, though the number varies depending on the acquisition choices. The pulse sequence, or the software that runs the radiofrequency antenna and magnetic coils which acquire the signal, determines how researchers acquire data. The repetition time between volumes, or TR, varies quite a bit across studies but typical values for a whole-brain acquisition have historically been about 2-3 seconds. However, recent imaging advances now make it possible to collect a whole brain volume in 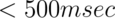 . Experiments can be brief at 6 minutes of functional time (e.g. 180 volumes), but experiments that include 40 or more minutes of functional time, with over 1,000 volumes measured at a typical TR, are not uncommon in practice.
. Experiments can be brief at 6 minutes of functional time (e.g. 180 volumes), but experiments that include 40 or more minutes of functional time, with over 1,000 volumes measured at a typical TR, are not uncommon in practice.
Thus fMRI data comprise hundreds to thousands of images in a time series. As local regions’ oxygen metabolism and blood flow change, researchers use fluctuations in the measured signal to make inferences about brain activity and connectivity. The usual approach towards assessing brain activity is based on examining average fluctuations locked to particular experimental conditions or events. We refer to this as task-based fMRI. Researchers assess brain connectivity by examining associations in the fluctuations among voxels with or without task condition influence analyses.
A simple, canonical example of a task-based fMRI experiment is a motor task. LetÕs say we want to examine activity increases in the motor cortex when participants execute simple finger movements. Researchers often use such tasks as quality control assessments in order to check signal and analysis quality. Participants might alternate between 20-second long blocks of finger tapping and 20 seconds of rest. This is a `block design’, illustrated in Figure 6.6’s bottom panel. Not all designs are equally efficient or powerful, but 20-second long blocks have good properties in particular; we will return to this concept in later chapters.
Statistical analysis
Once we have run an experiment and say obtained motor task data for a group of participants, we are ready to analyze the data. Recall that the fMRI dataset contains a time series of each voxel’s signal values. A basic analysis will study each participant’s data one person and one voxel at a time. fMRI data are quite noisy, so we use statistical analysis to determine whether a signal change is consistently associated with the finger-tapping task.
The first step of statistical analysis is to fit a model to each voxel’s time series. In this case, the model simply states that activity levels are different between finger-tapping and control periods. We can use a t-test to examine how large the difference (or `contrast’) is between finger tapping and rest divided by the noise measure (i.e. error variability). Then we make a map of the resulting voxel t-values and their associated p-values, which provides evidence to evaluate the null hypothesis of no task effect. Again, each voxel corresponds to a spatial location and has an associated statistic that represents the evidential strength for task-related effects. Researchers usually threshold these maps by applying a statistical cutoff related to the p-value, so scientific papers only plot and discuss voxels with sufficient evidence of an effect.
The description above covers the basics of a simple statistical analysis. However, it leaves out one key detail. fMRI activity, either BOLD or ASL, does not rise instantaneously when the task begins. Rather it increases over several seconds as blood flow function increases, peaks at about 5-6 seconds after the increase in local brain metabolic demand and decreases after 10-15 seconds. This function over time is the hemodynamic response function or the HRF which researchers must measure, or else assume a canonical function, to be able to perform a reasonably accurate analysis. Figure 7.7 shows a canonical HRF widely used as a model for fMRI responses. One piece of good news is that even very brief neural events (e.g. a 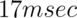 stimulus presentation) can reliably elicit measurable hemodyamic responses, so fMRI can be sensitive to short events. Another is that even with a complex neural event series or sustained blocks as in our finger-tapping experiment, we can still account for the HRF in our analysis.
stimulus presentation) can reliably elicit measurable hemodyamic responses, so fMRI can be sensitive to short events. Another is that even with a complex neural event series or sustained blocks as in our finger-tapping experiment, we can still account for the HRF in our analysis.

The description above highlights the fact that fMRI data analysis is fundamentally a time series problem. However, it’s a time series problem on steroids because every voxel has its own time series and there are about 100,000 voxels. The concept of analyzing voxels individually is sometimes called mass univariate analysis; in this, we treat all of the voxels separately then construct a map of the statistical results at each voxel. Other techniques that do not separate the voxels are becoming more widely used. We typically refer to these techniques as multivariate analyses because they are multivariate in brain space and model multiple voxels simultaneously.
Clearly fMRI data analysis is a massive data problem. Each brain volume consists of roughly 100,000 different voxel measurements. Each experiment might contain 1,000 brain volumes or more. And we might repeat each experiment for multiple subjects, maybe 20, 30, or 40, but sometimes hundreds or thousands, to facilitate population inference, i.e. making generalizable conclusions about human brain function. Because of both the amount of data and its complexity, fMRI data statistical analysis is challenging. The signal of interest is relatively weak and the data exhibits a complicated temporal and spatial noise structure. Thus there are ample opportunities to develop new increasingly sophisticated and powerful statistical techniques.
Data structure in fMRI experiments
Hierarchical data structure
fMRI data has a hierarchical structure, as Figure 7.8 shows. Understanding this structure and dealing with it appropriately is important when undertaking fMRI data preprocessing and statistical analysis.

The vast majority of experiments include many different participants’ data. This is critical in order to obtain population generalizable results - i.e. results that are not just idiosyncratic features of the individuals we happened to study but rather that constitute general conclusions that apply to new individuals. Each participant (sometimes called `subject’) performs the same task or tasks. Sometimes we nest participants within groups, such as patient group versus controls or elderly individuals versus young. In other cases, there is just one group and researchers’ interests are in studying experimental manipulations, behaviors, or other outcomes measured within-person. Even if we do not organize participants into groups, it is still possible relate brain activity differences to individual person-level variables (e.g. age, performance, or other variables).
Experiments entail collecting many repeated measurements on each participant over time. We may scan each participant longitudinally in multiple sessions. During a session, it is typical to start and stop the scanner multiple times, collecting data for brief periods - usually 4-10 minutes. We refer to these as runs. Because head movement during the scans is particularly problematic, short runs are advisable to give participants a break and allow them to communicate with the experimenters if necessary (though speaking can induce additional head movement!). Each run, in turn, entails a series of brain volumes, one per TR, nested within task conditions (e.g. finger tapping and rest). Each volume consists of multiple slices acquired sequentially, and each slice contains many voxels.
Often we analyze each participant separately with a mass univariate analysis of each voxel’s experimental effects. This is a first level analysis. The resulting maps of experimental effect magnitudes, called contrast maps, become the data for a second level analysis of effect reliability across participants, which includes differences between groups and effects of individual differences.
Image file formats
One barrier to entry in fMRI analysis is that the image data are not simple text files. Rather they are stored in specific customized formats along with associated `meta-data’ or information about the imaging parameters.
Historically, data formats differed widely across scanner manufacturers and software packages. For example, raw data on General Electric scanners are in a proprietary format called `P-files’ and on Siemens scanners as DICOM files (which stands for Digital Imaging and Communications in Medicine, http://www.dicomlibrary.com/dicom/).
DICOM files contain a single slice’s data at a single time point, with extensive `header’ information, though different scanners use and store this differently. A study can thus include millions of files, which presents logistical challenges with many file management systems. Therefore preprocessing and statistical analysis packages usually require that we convert these images into other, standard formats.
Though these standard formats also differed across packages, most now have the facility to read and write NIfTI (which stands for Neuroimaging Informatics Technology Initiative) images, a standard 3-D or 4-D file format. A 3-D NIfTi file contains one image per volume, while a 4-D NIfTi file often contains a single person’s time series of image volumes. Both these files have a .nii extension. A related older file format less standardly used across software packages is the Analyze image format which has .img extensions and associated separate header files containing meta-data with .hdr extensions.
It is important to exercise caution when reading and writing files across various software packages because these packages use the meta-data differently. Some researchers thus feel uncomfortable about mixing and matching algorithms from different software packages, though it can be done if one is cautious and meticulous.
One of the biggest issues of which to be aware relates to flipping images in the 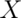 direction, from left to right. The brain is largely symmetrical, which makes it difficult to tell if an image display’s left side, such as in Figure 7.9, is on the left or right side of the brain. The fact that there are two orientations typically used to view images further complicates this point of potential confusion. If the image display is in radiological format, the brain’s left side is on the displayed image’s right side. This display is as though one looks up at a person’s brain from their feet. If images are in neurological format, the brain’s right side is on the displayed image’s right side. This is the format most cognitive neuroscience research uses. Though imaging software should keep track of format, different packages use header information related to flipping differently and custom reconstruction and stacking code at different research centers can also treat the image orientation information differently. As a result many, many errors have undoubtedly occurred.
direction, from left to right. The brain is largely symmetrical, which makes it difficult to tell if an image display’s left side, such as in Figure 7.9, is on the left or right side of the brain. The fact that there are two orientations typically used to view images further complicates this point of potential confusion. If the image display is in radiological format, the brain’s left side is on the displayed image’s right side. This display is as though one looks up at a person’s brain from their feet. If images are in neurological format, the brain’s right side is on the displayed image’s right side. This is the format most cognitive neuroscience research uses. Though imaging software should keep track of format, different packages use header information related to flipping differently and custom reconstruction and stacking code at different research centers can also treat the image orientation information differently. As a result many, many errors have undoubtedly occurred.

A number of strategies can help avoid this error. Using NIfTi format helps, as does consistent use of a single software package. Researchers often tape a Vitamin E capsule to the same side of each participant’s head. This produces a bright spot on the image in one hemisphere. Finally, we can heuristically check the flipping by viewing the images, because the left occipital lobe’s larger size in most people causes the calcarine fissure’s deviation to the right as it courses from front to back. This asymmetry is prominent in the structural image shown in Figure 6.9.
Conclusions
In this chapter, we briefly covered the major steps in fMRI data processing and analysis and some of the most commonly used terminology. In addition, we reviewed the hierarchical structure of fMRI data and common image file formats. In later chapters, we will cover the design, acquisition, preprocessing, and statistical analysis of fMRI data in more detail.