Chapter 1 — Generative AI: A New Frontier for Scientific Discovery
The New Frontier of Scientific Discovery
We stand at an extraordinary moment in the history of science. For centuries, the pace of discovery has been limited by human capacity—our ability to read literature, design experiments, analyze data, and synthesize knowledge. But a fundamental shift is underway. Generative Artificial Intelligence (AI) is not merely automating workflows; it is expanding what is scientifically possible [1].
Just as the microscope, the computer, and genome sequencing each transformed how we perceive nature, Generative AI now transforms our ability to understand and create scientific knowledge. AI systems design novel antibiotics by exploring chemical spaces that would take humans millennia to search [2, 3]. They predict protein structures with atomic precision—work recognized with the 2024 Nobel Prize in Chemistry [4, 5]. AI co-scientists generate hypotheses from vast textual corpora in days that previously required years of iterative research [6, 7], identify patterns in climate and Earth system data that are difficult to capture with traditional methods [8–10], and propose entirely new materials and reaction pathways [3, 11].
This is not science fiction—this is the reality of Generative AI today [1].
The AI Revolution in Scientific Discovery
From Analysis to Generation
Scientific computing has evolved through clear stages:
| Era | Core Capability | Tools | Outcome |
|---|---|---|---|
| 1960s–1980s | Digitization | Databases | Data preservation |
| 1990s–2010s | Statistical & ML methods | Regression, SVMs, random forests | Pattern recognition |
| 2020s– | Generative AI | Transformers, diffusion models | Creation of new data & hypotheses |
Traditional scientific discovery has often followed a cycle of observe → hypothesize → experiment → analyze → repeat [12]. Generative AI compresses and accelerates this loop [1, 7]:
- Literature review: Synthesizes insights from millions of papers [6, 7].
- Hypothesis generation: Identifies non-obvious links across fields, with AI co-scientists reducing hypothesis generation from weeks to days [1, 7].
- Experimental design: Proposes thousands of candidate molecules or materials for testing [2, 3, 11].
- Data analysis: Finds structure in high-dimensional scientific datasets [8–10, 12].
The Acceleration Effect
The effect is both quantitative and qualitative [1]:
- Speed: AI approaches can shorten drug discovery timelines from ~10–15 years to ~3–5 in some workflows, with AI-designed drugs showing 80–90% success rates in Phase I trials compared to traditional 40–65% [2, 13].
- Scale: Virtual screening has been used to evaluate extensive compound libraries in silico for drug discovery, supporting efficient prioritization of candidates[3].
- Efficiency: Weather forecasts that once required hours on supercomputers can be generated in minutes with AI models like GenCast [9, 14].
More profoundly, AI enables scientists to explore vast combinatorial spaces and ask what if? at scales that would be infeasible with manual or purely equation-based approaches [1, 12].
What Makes Generative AI Different from Traditional ML?
Traditional Machine Learning — Pattern Recognition
Traditional machine learning (ML) excels at discriminative tasks such as classification, regression, and clustering—learning mappings from inputs to outputs and identifying patterns in existing data [12].
Generative AI — Creating the New
Generative models go further: they learn the distribution underlying the data and can sample from it to create new, realistic instances [15, 16]. This enables capabilities such as:
| Capability | Example | Outcome |
|---|---|---|
| Generation | Design a new molecule with desired properties | Novel compounds [2, 3] |
| Completion | Fill missing regions in a protein structure or sequence | Plausible biological variants [4, 5] |
| Translation | Text description → molecule, code, or figure | Cross-modal synthesis [1, 6] |
| Synthesis | Combine text, code, and data | New simulations, analyses, or visualizations [6] |
Core Technologies Powering Generative AI
1. Transformers and Large Language Models (LLMs)
Transformers introduced an attention-based architecture that models long-range dependencies and contextual relationships in sequences [15]. Large Language Models (LLMs) built on this architecture can:
- Summarize and synthesize scientific literature
- Generate runnable analysis code and scripts
- Act as domain-aware assistants that interact with text, equations, and data [1, 6]
- Serve as AI co-scientists that generate novel hypotheses through multi-agent debate and evolution [7]
2. Diffusion Models
Diffusion models learn to reverse a gradual noising process to generate high-fidelity samples [16–18]. They are increasingly used for:
- Molecular and materials design [3, 11]
- Protein and structural biology tasks, including AlphaFold 3’s prediction of biomolecular interactions [4, 5]
- Probabilistic weather forecasting with models like GenCast [9, 14]
3. Variational Autoencoders (VAEs) and Generative Adversarial Networks (GANs)
Variational Autoencoders (VAEs) and Generative Adversarial Networks (GANs) provide earlier but still important frameworks for generative modeling [19, 20]. They support:
- Learning low-dimensional latent representations of complex systems
- Creating synthetic datasets to augment limited experimental data
- Constructing surrogate models for expensive simulations
The Pre-Training Revolution
Modern generative AI thrives on pre-training: models are first trained on large, generic corpora (text, code, images, or scientific data) and then fine-tuned on specific tasks or domains [6]. This strategy is particularly valuable in science, where high-quality labeled data are often scarce or costly to obtain.
A language model pre-trained on broad text can be adapted to domains such as biomedicine, materials science, or geosciences. Protein models pre-trained on millions of sequences can specialize in a narrow protein family with comparatively small task-specific datasets [4, 5].
Generative AI Across Scientific Disciplines
Physical and Chemical Sciences
In the physical and chemical sciences, generative AI is already reshaping how we search, design, and reason about complex systems:
- Materials Design: Exploring enormous chemical spaces to propose new superconductors, catalysts, or battery materials with targeted properties [3, 11].
- Reaction and Synthesis Planning: Suggesting reaction products and retrosynthetic routes, accelerating the path from concept to synthesis [3].
- Mathematical Reasoning: AlphaProof and AlphaGeometry 2 achieved silver-medal level performance at the 2024 International Mathematical Olympiad, and an advanced Gemini model achieved gold-medal standard in 2025 [21, 22].
Chemistry and Drug Discovery
In chemistry and drug discovery, generative models help to:
- Propose novel antibiotics and therapeutics with desired efficacy and safety profiles [2].
- Optimize molecules for multiple objectives (binding, solubility, toxicity, synthesizability) [3, 11].
- Accelerate clinical progress: AI-assisted drug candidates achieved Phase I success rates of nearly 90%, compared with industry averages of 40–65% [13].
Life and Environmental Sciences
- Protein Engineering: AlphaFold 3 predicts the structure and interactions of proteins, DNA, RNA, ligands, and ions with unprecedented accuracy, achieving at least 50% improvement over existing methods [5].
- Genomics and Systems Biology: Modeling sequence–function relationships and generating candidate variants for experimental follow-up [1].
- Medical AI: Supporting risk prediction, trial design, and treatment-effect modeling when combined with causal and statistical frameworks [1, 12].
Climate and Earth System Science
- Weather Forecasting: GenCast, a diffusion-based probabilistic model, outperforms the world’s leading operational forecast (ECMWF ENS) in 97% of scenarios, delivering 15-day forecasts in 8 minutes [9, 14].
- Earth System Foundations: Large foundation models that integrate atmosphere, ocean, land, and cryosphere processes, including Google’s Aurora and WeatherNext models [10].
- Surrogate Modeling: Physics-informed and data-driven surrogates that emulate expensive numerical models at a fraction of the computational cost [8–10, 23].
Mathematical Foundations and Methods
Generative AI is also driving progress in the mathematics and methodology of scientific machine learning:
- Data-Driven Dynamical Systems: Learning governing dynamics and control laws directly from measurements or simulations [12].
- Physics-Informed Neural Networks (PINNs): Incorporating partial differential equations and other physical constraints directly into neural network training [24, 25].
- Uncertainty Quantification: Developing methods to quantify confidence and propagate uncertainty through AI-based scientific pipelines [25].
- Formal Mathematical Reasoning: AlphaProof combines reinforcement learning with formal theorem proving in Lean to generate verifiable mathematical proofs [21].
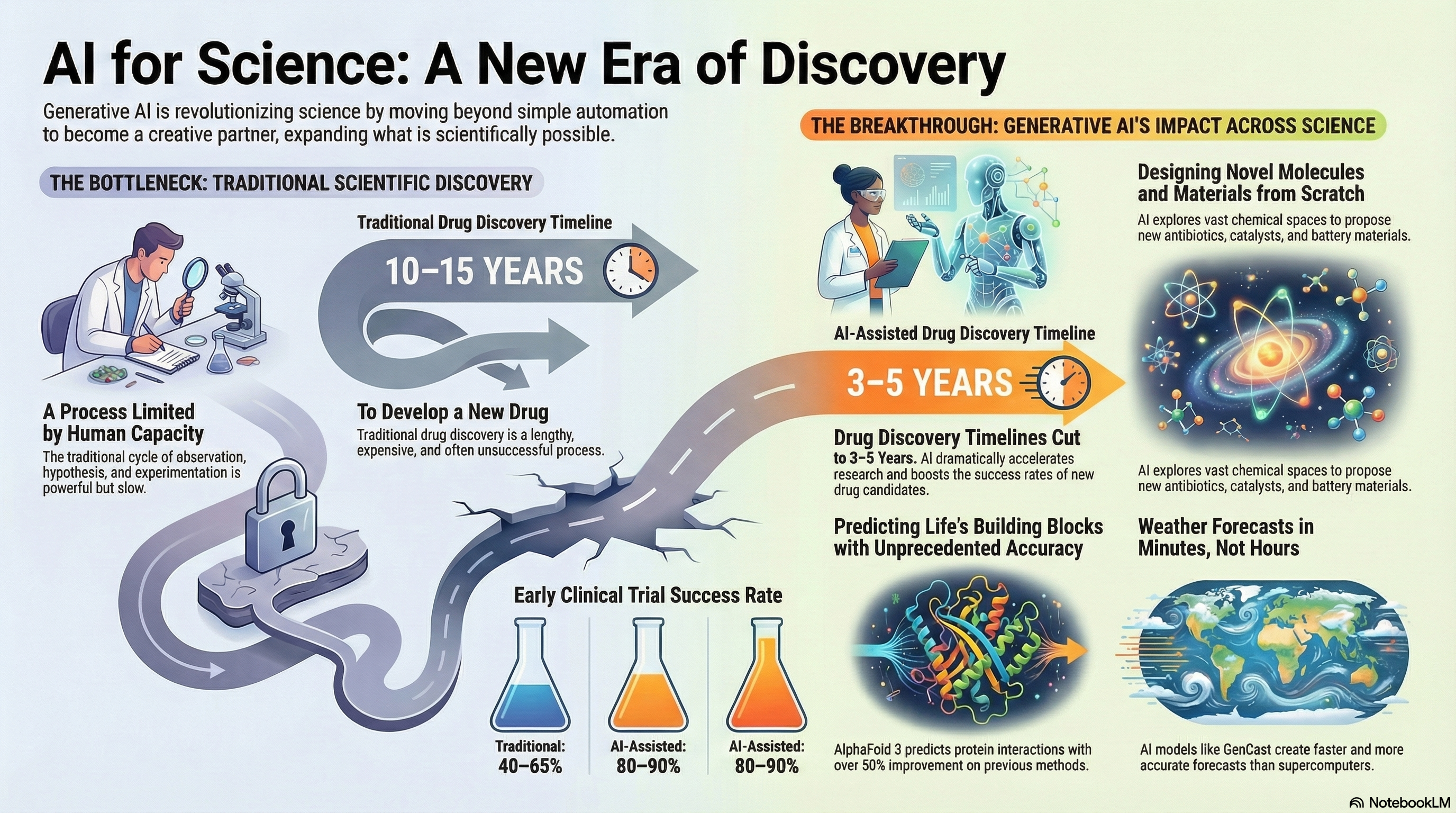
Figure 1.1. AI for Science: A New Era of Discovery. Traditional scientific discovery (left) is limited by human capacity, with drug development taking 10–15 years and early clinical trial success rates of 40–65%. Generative AI (right) is transforming this landscape by reducing drug discovery timelines to 3–5 years, improving clinical trial success rates to 80–90%, enabling novel molecule and materials design, predicting protein structures with unprecedented accuracy (AlphaFold 3), and accelerating weather forecasts from hours to minutes (GenCast). (Created by Google NotebookLM)
Cross-Cutting Capabilities
Several techniques and ideas cut across scientific domains:
- Physics-Informed Models: Embedding conservation laws, symmetries, and PDEs in neural architectures to ensure physically consistent predictions [24, 25].
- Latent-Variable and Generative Representations: Using VAEs, GANs, and diffusion models to learn compact representations of high-dimensional systems [16–20].
- Multimodal Integration: Combining text, images, time series, and spatial fields to build holistic models of complex phenomena [1, 6].
- Multi-Agent AI Systems: Frameworks like Google’s AI co-scientist use specialized agents for hypothesis generation, debate, and evolution [7].
A New Scientific Partner
Generative AI is not replacing scientists—it is becoming a new kind of collaborator [1]. It can:
- Expand the hypothesis space far beyond what any individual or team could enumerate
- Accelerate cycles of modeling, simulation, and experiment
- Enable researchers with limited computational or institutional resources to use state-of-the-art tools
- Bridge previously separate fields by revealing shared patterns in their data and models
- Recapitulate discoveries in days that previously required years of iterative research [7]
The most impactful discoveries are likely to emerge where human insight and machine-generated ideas interact in a tight loop.
The Path Forward
The rest of this book will help you:
- Understand the core architectures (transformers, diffusion models, scientific ML) [15–18, 24, 25].
- Adapt and fine-tune models to your own scientific domain [2–5, 11].
- Evaluate and validate models with appropriate statistical and physical checks [12, 23, 25].
- Design human–AI workflows in which AI accelerates, rather than distorts, the scientific process [1].
The revolution is already underway. In my view, scientists who understand and apply generative AI will stand at the forefront of discovery in every field [1].
References and Further Readings
Wang, H., Fu, T., Du, Y., Gao, W., Huang, K., Liu, Z., et al. (2023). Scientific discovery in the age of artificial intelligence. Nature, 620(7972), 47–60. https://doi.org/10.1038/s41586-023-06221-2
Stokes, J. M., Yang, K., Swanson, K., Jin, W., Cubillos-Ruiz, A., Donghia, N. M., et al. (2020). A deep learning approach to antibiotic discovery. Cell, 180(4), 688–702. https://doi.org/10.1016/j.cell.2020.01.021
Gómez-Bombarelli, R., Wei, J. N., Duvenaud, D., Hernández-Lobato, J. M., Sánchez-Lengeling, B., Sheberla, D., et al. (2018). Automatic chemical design using a data-driven continuous representation of molecules. ACS Central Science, 4(2), 268–276. https://doi.org/10.1021/acscentsci.7b00572
Jumper, J., Evans, R., Pritzel, A., Green, T., Figurnov, M., Ronneberger, O., et al. (2021). Highly accurate protein structure prediction with AlphaFold. Nature, 596(7873), 583–589. https://doi.org/10.1038/s41586-021-03819-2
Abramson, J., Adler, J., Dunger, J., Evans, R., Green, T., Pritzel, A., et al. (2024). Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature, 630(8016), 493–500. https://doi.org/10.1038/s41586-024-07487-w
Brown, T., Mann, B., Ryder, N., Subbiah, M., Kaplan, J. D., Dhariwal, P., et al. (2020). Language models are few-shot learners. Advances in Neural Information Processing Systems, 33, 1877–1901. https://arxiv.org/abs/2005.14165
Gottweis, J., Weng, W.-H., Daryin, A., Liu, Y., Ceze, L., Natarajan, V., et al. (2025). Towards an AI co-scientist. arXiv preprint arXiv:2502.18864. https://arxiv.org/abs/2502.18864
Reichstein, M., Camps-Valls, G., Stevens, B., Jung, M., Denzler, J., Carvalhais, N., & Prabhat. (2019). Deep learning and process understanding for data-driven Earth system science. Nature, 566(7743), 195–204. https://doi.org/10.1038/s41586-019-0912-1
Price, I., Sanchez-Gonzalez, A., Alet, F., Andersson, T. R., El-Kadi, A., Masters, D., et al. (2024). Probabilistic weather forecasting with machine learning. Nature, 636, 84–90. https://doi.org/10.1038/s41586-024-08252-9
Bodnar, C., Bruinsma, W. P., Lucic, A., Stanley, M., Allen, A., Brandstetter, J., et al. (2024). Aurora: A foundation model of the atmosphere. arXiv preprint arXiv:2405.13063. https://arxiv.org/abs/2405.13063
Angello, N. H., Friday, D. M., Hwang, C., Yi, S., Cheng, A. H., Torres-Flores, T. C., et al. (2024). Closed-loop transfer enables artificial intelligence to yield chemical knowledge. Nature, 633(8029), 351–358. https://doi.org/10.1038/s41586-024-07021-y
Brunton, S. L., & Kutz, J. N. (2022). Data-driven science and engineering: Machine learning, dynamical systems, and control (2nd ed.). Cambridge University Press. https://doi.org/10.1017/9781108380690
AI in Drug Discovery (2024). Industry analysis reports indicate AI-assisted drug candidates achieving Phase I success rates of nearly 90%, compared with industry averages of 40–65%. Drug Discovery News, October 2025. https://pubmed.ncbi.nlm.nih.gov/38692505
Google DeepMind (2024). GenCast predicts weather and the risks of extreme conditions with state-of-the-art accuracy. https://deepmind.google/blog/gencast-predicts-weather-and-the-risks-of-extreme-conditions-with-sota-accuracy/
Vaswani, A., Shazeer, N., Parmar, N., Uszkoreit, J., Jones, L., Gomez, A. N., et al. (2017). Attention is all you need. Advances in Neural Information Processing Systems, 30. https://arxiv.org/abs/1706.03762
Ho, J., Jain, A., & Abbeel, P. (2020). Denoising diffusion probabilistic models. Advances in Neural Information Processing Systems, 33, 6840–6851. https://arxiv.org/abs/2006.11239
Dhariwal, P., & Nichol, A. (2021). Diffusion models beat GANs on image synthesis. Advances in Neural Information Processing Systems, 34, 8780–8794. https://arxiv.org/abs/2105.05233
Song, Y., Sohl-Dickstein, J., Kingma, D. P., Kumar, A., Ermon, S., & Poole, B. (2021). Score-based generative modeling through stochastic differential equations. Proceedings of ICLR. https://arxiv.org/abs/2011.13456
Kingma, D. P., & Welling, M. (2014). Auto-encoding variational Bayes. Proceedings of ICLR. https://arxiv.org/abs/1312.6114
Goodfellow, I., Pouget-Abadie, J., Mirza, M., Xu, B., Warde-Farley, D., Ozair, S., et al. (2014). Generative adversarial networks. Advances in Neural Information Processing Systems, 27. https://arxiv.org/abs/1406.2661
Google DeepMind (2024). AI achieves silver-medal standard solving International Mathematical Olympiad problems. https://deepmind.google/blog/ai-solves-imo-problems-at-silver-medal-level/
Google DeepMind (2025). Advanced version of Gemini with Deep Think officially achieves gold-medal standard at the International Mathematical Olympiad. https://deepmind.google/blog/advanced-version-of-gemini-with-deep-think-officially-achieves-gold-medal-standard-at-the-international-mathematical-olympiad/
Carleo, G., Cirac, I., Cranmer, K., Daudet, L., Schuld, M., Tishby, N., et al. (2019). Machine learning and the physical sciences. Reviews of Modern Physics, 91(4), 045002. https://doi.org/10.1103/RevModPhys.91.045002
Raissi, M., Perdikaris, P., & Karniadakis, G. E. (2019). Physics-informed neural networks: A deep learning framework for solving forward and inverse problems involving nonlinear partial differential equations. Journal of Computational Physics, 378, 686–707. https://doi.org/10.1016/j.jcp.2018.10.045
Karniadakis, G. E., Kevrekidis, I. G., Lu, L., Perdikaris, P., Wang, S., & Yang, L. (2021). Physics-informed machine learning. Nature Reviews Physics, 3(6), 422–440. https://doi.org/10.1038/s42254-021-00314-5
Next → Chapter 2: introduces the core architectures powering generative AI.